Data Harmonization in Healthcare: Building the Foundation for AI-Ready Clinical Data
We can’t talk seriously about AI in healthcare without understanding the data infrastructure behind it. From HL7 v2 to FHIR, SNOMED, and MCP, I break down how clinical data actually moves and why every healthcare leader needs to grasp this before approving the next AI pilot.
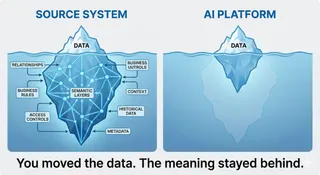
The Problem: A Landscape of Fragmented Data
If you've spent any time working with healthcare data, you know the reality on the ground: clinical information is scattered across dozens of disconnected systems, each speaking its own dialect. An EHR stores diagnoses in HL7 v2 or C-CDA documents. The radiology department transmits imaging metadata via DICOM. Lab systems use their own internal codes. Billing runs on claims data with ICD-10 codes that were chosen more for reimbursement than clinical accuracy (e.g. W61.62XA – Struck by duck, initial encounter).
None of these systems were designed to talk to each other. They were built over decades by different vendors, for different purposes, and they represent the same underlying clinical reality in fundamentally different ways. One hospital's "Type 2 Diabetes Mellitus" might be stored as a SNOMED code in the EHR, an ICD-10 code on the claim, a free-text note in a lab order, and a local shorthand in the pharmacy system, all describing the same condition for the same patient.
This is what we mean by fragmented data, and it's the central obstacle standing between healthcare organizations and the AI-driven future they're investing in. You can't build reliable predictive models, clinical decision support, or agentic AI workflows on a foundation of inconsistent, ambiguously coded information.
Data, Semantics, and Ontology: Three Layers of Meaning
Before diving into specific standards, it helps to understand why data harmonization demands so much rigor in the first place, and it starts with a fundamental difference between how humans and machines process information.
The human brain is an association engine. We constantly make connections between what we see, hear, and experience, drawing on context without conscious effort. A doctor reading a chart that says "sugar is high" immediately understands this likely refers to blood glucose, connects it to a possible diabetes diagnosis, considers medication implications, and thinks about dietary factors, all in a fraction of a second. We fill in gaps, resolve ambiguity, and reason across loosely connected information effortlessly.
Software does not work this way. A machine has no intuition, no background experience, no ability to "just know" what a value probably means. If a data field contains "120" with no label, no unit, and no code system, a computer cannot guess whether it's a heart rate, a glucose reading, or a systolic blood pressure. If two hospitals call the same lab test by different names, a machine sees two unrelated concepts. Unless something is defined precisely, with explicit structure, unambiguous codes, and formal relationships, software cannot interpret it at all.
This is the core tension in healthcare data. Clinicians operate in a world of rich context and implicit understanding. Machines require everything to be spelled out. The entire discipline of data harmonization exists to bridge that gap, to translate the messy, contextual, human world of clinical practice into something a computer can reliably process and reason about.
That bridge rests on three layers, and getting them right is where most AI and analytics projects in healthcare succeed or fail.
Data is a value within a structure. The number '120' by itself is just characters on a screen. Is it a heart rate or the patient weight? Data without context is ambiguous. It's only when '120' sits inside a field like 'systolic_bp' in a clinical table, with a system identifier, a unit, and a timestamp, that it begins to become useful. A string like "E11.9" means nothing until you know it lives in a 'diagnosis_code' field and belongs to the ICD-10 code system, version 2019. Data, in other words, is raw values plus the schema that gives them a minimum of structure.
Semantics is what the data means, and it goes well beyond mapping a code to a human-readable label. Take LOINC code '718-7'. Semantics tells us this code represents Hemoglobin [Mass/volume] in Blood. But it also tells us that this is a quantitative lab test, that it uses specific units (g/dL), that it belongs to the hematology domain, and that it has known synonyms across systems. Semantics encompasses code system definitions, contextual meaning, units, synonyms, and versioning. It is what ensures that when two different hospitals send a hemoglobin result, a consuming system can recognize them as the same clinical concept, even if the source systems represent them differently.
Ontology goes further still. Where semantics labels and defines individual concepts, an ontology formalizes the logical relationships between them, enabling machines to reason and infer. Consider SNOMED CT's representation of pneumonia. It doesn't just say "Pneumonia is code X." It defines that Pneumonia is-a Infectious disease, that its finding site is Lung structure, and that its causative agent may be Bacteria. These aren't just hierarchical parent-child links. A true ontology includes is-a relationships, part-of relationships, causal and temporal relationships, and formal logical constraints. This richness is what allows a reasoning engine to infer that if a disease affects the lung, it relates to respiratory function, and if it's bacterial in origin, antibiotics may be relevant, without any of that being explicitly programmed.
The key differentiator: a terminology labels concepts, an ontology formalizes logic between them.
Clarifying the common confusion. In healthcare, the terms "ontology," "terminology," and "data model" are used interchangeably, but they shouldn't be. A terminology system (ICD-10, LOINC) provides vocabulary. An ontology (SNOMED CT is the closest example in clinical practice) provides logical structure that enables inference. A data model (FHIR, OMOP) defines how data is stored and exchanged. FHIR is not an ontology. ICD-10 is not a true ontology. Getting this distinction right is the first step toward a harmonization strategy that actually holds up, because a machine will never figure out the difference on its own. It needs us to get it right at the foundation.
The Big Three: FHIR, OMOP, and SNOMED CT
Healthcare has converged on a handful of major standards, each designed for a specific purpose. They're complementary, not competing.
FHIR - The Exchange Standard
FHIR (Fast Healthcare Interoperability Resources), developed by HL7, is the modern standard for moving data between systems. Think of it as the universal packaging format for clinical information.
FHIR breaks clinical data into discrete Resources: Patient, Observation, Condition, MedicationRequest, and so on, formatted as JSON or XML. Its API-first, RESTful design makes it the natural fit for real-time clinical applications, patient-facing tools, and increasingly, AI agents.
Critically, FHIR is a carrier, not an ontology. It doesn't define what "Diabetes" is. Instead, it provides a structured slot where you insert a code from SNOMED CT or ICD-10. FHIR's power is in its structure: it gives you discrete, queryable, machine-readable building blocks rather than monolithic documents. An AI agent can request a specific Observation resource for a patient's latest HbA1c result instead of parsing a 50-page discharge summary.
FHIR also supports Profiles and Extensions, defined through Implementation Guides like US Core, that let you enforce strict data quality constraints on your canonical model, ensuring downstream consumers, whether human or AI, receive predictable, well-structured data.
OMOP - The Research Standard
The OMOP Common Data Model, maintained by the OHDSI community, was designed for a completely different use case: large-scale observational research and population health analytics.
OMOP standardizes disparate hospital databases into a single relational schema with tables like 'condition_occurrence', 'drug_exposure', and 'measurement'. Its key strength is that it maps all the local, "messy" codes from individual institutions into Standard Concepts, typically drawn from SNOMED CT and RxNorm, so that a research query written in London can execute identically against data in New York, Tokyo, or São Paulo.
You don't use OMOP to treat a patient in real-time. You use it to study 10,000 patients with the same condition, to train predictive models, or to run the kind of population-level SQL analytics that would be impractical against a live FHIR server.
SNOMED CT - The Clinical Ontology
SNOMED CT is the most comprehensive, multilingual clinical terminology in the world, with over 350,000 concepts connected by formal logical relationships. Unlike FHIR (a structural framework) or OMOP (a data model), SNOMED CT is the closest thing healthcare has to a true ontology.
Its poly-hierarchical structure means a single concept can have multiple parent relationships and multiple relationship types. "Diabetic retinopathy," for instance, is simultaneously a disorder of the eye and a complication of diabetes. This richness enables the kind of granularity clinicians need, not just "Fracture" but "Comminuted fracture of the left femur," and the kind of semantic reasoning AI systems need to draw inferences across a patient's clinical history.
SNOMED CT is widely used for clinical problem lists and findings and serves as a common semantic backbone for mapping and decision support, making it a critical layer that both FHIR and OMOP depend on.
The Supporting Cast: LOINC, RxNorm, ICD, and DICOM
No single standard covers everything. Several other terminologies and protocols fill essential gaps:
LOINC (Logical Observation Identifiers Names and Codes) is the universal language for lab results and clinical observations. When your AI agent needs to find a patient's blood glucose level, it should look for the appropriate LOINC glucose concept, for example 2345-7 (Glucose in Serum/Plasma) or 2339-0 (Glucose in Blood), not a hospital-specific string like "GLU-SERUM" or "glucose, random."
RxNorm provides standardized names for clinical drugs and dosages. It's what prevents an AI from treating "Lipitor 20mg" and "Atorvastatin Calcium 20mg" as two different medications. For medication reconciliation and drug interaction checking, RxNorm is indispensable.
ICD-10/11 is primarily a classification system used for billing and international mortality/morbidity reporting. It's less granular than SNOMED CT but ubiquitous in claims data, making it an important part of any ingestion pipeline.
DICOM is the global standard for medical imaging, X-rays, MRIs, CT scans, and their associated metadata. If your AI pipeline includes imaging data, DICOM is how it gets in the door.
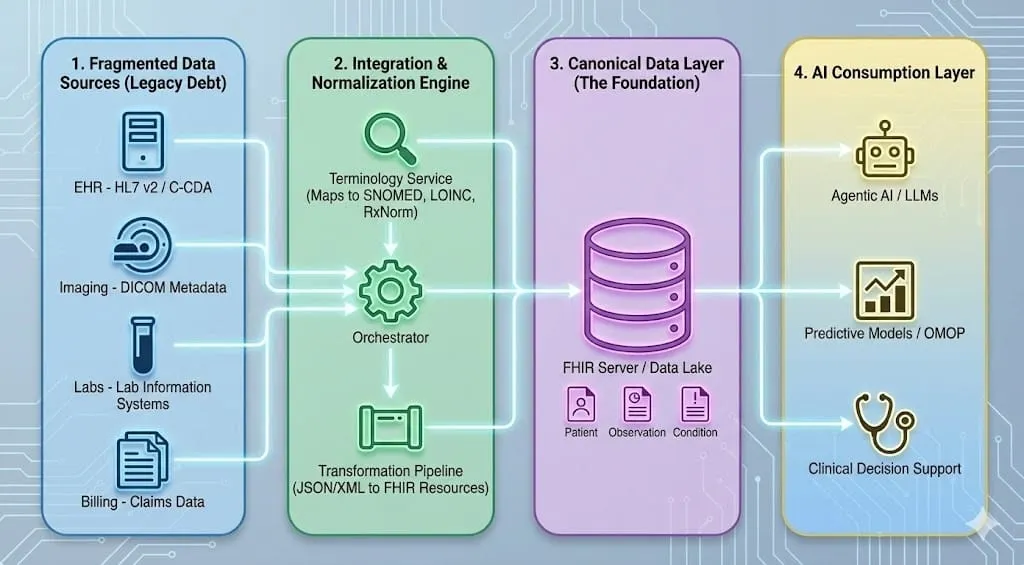
How They All Work Together
Picture a typical clinical workflow: a doctor records a diagnosis in an EHR using a SNOMED CT code which is then mapped to ICD-10 for billing purposes. The EHR sends that information to a specialist's office via a FHIR API. The associated lab results are coded in LOINC, the prescribed medication in RxNorm, and the chest X-ray is stored in DICOM format. Later, a researcher pulls all of this into an OMOP data warehouse to run a retrospective study across 50 hospitals.
Each standard has its lane. The art of data harmonization is getting them to work in concert.
Why This Matters for AI, and Not Only AI
It's worth pausing to make an important point: data harmonization is not a new idea driven by the rise of AI. These standards have been used for years, in some cases decades, to solve interoperability challenges across the healthcare technology landscape. Any solution built on clinical data benefits from harmonization. Analytics dashboards, quality reporting systems, clinical registries, patient portals, population health platforms, care coordination tools, all of them produce better outcomes when the underlying data is consistent, well-coded, and semantically clear. Organizations that invested in FHIR, SNOMED CT, and LOINC long before anyone was talking about large language models are now reaping the benefits, because they already have the clean, structured foundation that modern AI demands.
What AI does is raise the stakes. AI, whether it's a predictive model, a clinical decision support system, or an autonomous agent, is only as good as the data it consumes, and it is far less forgiving of ambiguity than a human analyst reviewing a dashboard.
The Data Level
An AI model trained on harmonized data, where every diagnosis maps to SNOMED, every lab result to LOINC, and every medication to RxNorm, can generalize across institutions, patient populations, and time periods. A model trained on raw, un-harmonized hospital data learns the idiosyncrasies of that one hospital's coding practices. It doesn't transfer. It doesn't scale.
FHIR's resource-based structure is particularly powerful here. Because each clinical concept is broken into a discrete, self-describing resource with explicit codes and references, an AI system can navigate a patient's record programmatically. It doesn't need to parse natural language or guess at the meaning of an ambiguous local code. The data describes itself.
The Connectivity Level: Enter MCP
But harmonized data sitting in a FHIR server is only half the picture. AI agents also need a standardized way to connect to and interact with that data, and this is where the Model Context Protocol (MCP) enters the picture.
MCP is an open protocol for connecting AI applications to external tools and data sources through a consistent interface, often described as a "USB-C" style connector for agentic workflows. In a healthcare context, an MCP server can sit alongside a FHIR API and present curated, task-oriented functions like "get most recent HbA1c," while still enforcing the same authentication, authorization, auditing, and least-privilege controls required for clinical systems.
This is a meaningful shift. Traditional system integration is about point-to-point API plumbing: each new consumer needs custom code to talk to each data source. MCP abstracts that away. An AI agent built with MCP support can interact with a FHIR API, an OMOP warehouse, or a DICOM archive through MCP-compatible tools, using the same protocol, without requiring custom integration logic for each data source..
An important caveat: MCP is emerging and not healthcare-specific, so it should be treated as an integration pattern that complements FHIR, not a replacement for interoperability standards or enterprise integration controls. In regulated environments, MCP endpoints need the same security posture as any clinical integration surface, strong authentication and authorization, audit trails, least-privilege tool scopes, and rigorous prompt-injection hardening. The protocol is promising, but it must be deployed with the same governance rigor as any other clinical data interface.
For clinical decision support and agentic AI workflows, the combination of harmonized data (FHIR + standard terminologies) and standardized connectivity (MCP) creates a foundation where AI agents can reliably reason across a patient's full clinical picture, labs, diagnoses, medications, imaging, without the brittle, custom integration work that has historically made healthcare AI so difficult to scale.
From Raw to Reasoning: A Lab Result's Journey
All of the standards and layers we've discussed can feel abstract, so let's walk through a single, concrete example: a patient's blood glucose test result, traveling from the moment it's generated in a hospital lab all the way to an AI agent that can reason about it.
Step 1: The Starting Point - A Raw HL7 Message
When a hospital lab system finishes processing a blood sample, it doesn't produce a nicely formatted document. It fires off an HL7 v2 message, a format designed in the 1980s that looks something like this:
OBX|1|NM|2345-7^Glucose^LN||126|mg/dL|70-100|H|||F
This is a positional format, meaning every piece of information sits in a specific slot separated by pipe characters. To a human reading it cold, it's barely intelligible. But buried in that string is everything we need: the test code (2345-7), the test name (Glucose), the code system (LN for LOINC), the result value (126), the unit (mg/dL), the normal range (70-100), a flag indicating the result is high (H), and a status saying the result is final (F).
The problem? This format is rigid, fragile, and carries almost no self-describing context. The field positions are defined by the HL7 v2 specification, not discoverable from the message itself. If you're a system that didn't already know the HL7 v2 segment layout by heart, you'd have no idea what field 5 or field 6 means. There's no label that says "this is the value" or "this is the unit." It's structure without semantics.
Step 2: Normalization - Making Sure "Glucose" Means Glucose
Before we can transform this into something richer, we need to verify and normalize the terminology. The message references LOINC code 2345-7, but not every source system is this cooperative. Another hospital's lab system might send the same glucose result with a local code like GLU-R or CHEM7-2, with no mention of LOINC at all.
This is where a terminology service steps in. It takes whatever code the source system provides and maps it to the correct standard vocabulary: LOINC 2345-7 for the test itself, and SNOMED CT or other relevant codes for any associated clinical context. But mapping codes is only part of what a terminology service does. It also validates whether codes are active or retired, normalizes units to the UCUM standard (the http://unitsofmeasure.org system you'll see in the FHIR resource below), manages version differences between code systems, expands value sets, and handles crosswalks between vocabularies, for example mapping a local code to LOINC, or a SNOMED concept to its ICD-10-CM equivalent for billing. The goal is simple: no matter how the source system chose to represent "blood glucose," by the time it leaves this step, it's expressed in a universal language that any downstream consumer can understand without guessing.
Step 3: Transformation - From Pipes to Self-Describing Data
Now comes the transformation into FHIR. That cryptic pipe-delimited string becomes a structured JSON resource that carries its meaning with it:
{
"resourceType": "Observation",
"status": "final",
"category": [{
"coding": [{
"system": "http://terminology.hl7.org/CodeSystem/observation-category",
"code": "laboratory",
"display": "Laboratory"
}]
}],
"code": {
"coding": [{
"system": "http://loinc.org",
"code": "2345-7",
"display": "Glucose [Mass/volume] in Serum or Plasma"
}]
},
"subject": {
"reference": "Patient/12345"
},
"valueQuantity": {
"value": 126,
"unit": "mg/dL",
"system": "http://unitsofmeasure.org",
"code": "mg/dL"
},
"referenceRange": [{
"low": { "value": 70, "unit": "mg/dL" },
"high": { "value": 100, "unit": "mg/dL" }
}],
"interpretation": [{
"coding": [{
"system": "http://terminology.hl7.org/CodeSystem/v3-ObservationInterpretation",
"code": "H",
"display": "High"
}]
}]
} Look at what's happened. Every piece of information that was hiding in a positional slot now has a name, a context, and a source. The code isn't just 2345-7, it comes with the system it belongs to (http://loinc.org) and a human-readable description. The value isn't just 126 floating in field 5, it's explicitly labeled as a quantity with a unit and a standardized unit code. The "H" flag isn't just a letter, it's linked to a formal interpretation code system that defines what "High" means.
This is what we mean by self-describing data. A system encountering this resource for the first time, with no prior knowledge of the source hospital, can understand exactly what it's looking at. The data carries its own context.
Step 4: The AI Agent - Reasoning Through MCP
Here's where it all pays off. An AI agent, say a clinical decision support tool reviewing a patient's chart, needs to assess whether this patient might be at risk for diabetes. In a traditional integration, someone would have had to write custom code teaching the agent how to call the specific hospital's API, parse its particular response format, and interpret its local codes.
With an MCP server sitting alongside the FHIR API, the agent can express what it needs through a structured, task-oriented interface:
"Get the most recent glucose result for Patient 12345"
The MCP server translates this into the appropriate FHIR query, retrieves the Observation resource, and returns it to the agent in a form it can immediately work with. The agent doesn't need to know anything about FHIR query syntax or REST endpoints. The MCP layer handles the plumbing, while the underlying FHIR server continues to enforce authentication, authorization, auditing, and consent controls
Now the agent has everything it needs to reason. The glucose value is 126 mg/dL. The reference range says normal is 70 to 100. The interpretation flag says "High." Because the data is coded in LOINC, the agent knows with certainty this is a serum glucose measurement, not some other test that happens to share a name. Because the value has explicit units, there's no ambiguity about scale. Because the reference range is included, the agent can assess severity without needing a separate lookup table.
The agent can now combine this result with other harmonized data in the patient's record, a recent HbA1c result (also LOINC-coded), a BMI observation, a family history of diabetes (SNOMED-coded), and reach a well-supported conclusion: this patient's glucose is elevated, their pattern of results is consistent with pre-diabetes, and a follow-up should be recommended.
None of that reasoning is possible if the data is still sitting in a pipe-delimited HL7 message with local codes that only one hospital's system understands.
That's the whole journey. A positional string generated by a 1980s-era lab system becomes a semantically rich, self-describing, machine-readable resource that an AI agent can access through a standardized protocol and reason about with confidence. Every layer of the harmonization pipeline, the terminology mapping, the FHIR transformation, the MCP connectivity, exists to make that final step possible.
Beyond Structure: Provenance, Trust, and Governance
Harmonizing the format and coding of clinical data is necessary, but it's not sufficient. In regulated healthcare environments, "AI-ready" data also needs to carry information about where it came from, how trustworthy it is, and who is allowed to see it. This is where harmonization meets governance, and it's often where conversations with compliance, legal, and clinical leadership get real.
Provenance means knowing who recorded a data element, when, from which source system, and through what process it arrived in your canonical layer. FHIR has a dedicated Provenance resource for exactly this purpose. Without it, an AI agent may be reasoning on data it has no basis to trust.
Data quality and completeness must be tracked explicitly. A harmonized record that is missing half its lab results or carries stale medication data can lead an AI agent to confident but wrong conclusions. Quality metadata, flagging completeness, timeliness, and validation status, should travel with the data, not live in a separate report no one reads.
Consent, segmentation, and disclosure govern what data can be used, by whom, and for what purpose. Behavioral health records, substance abuse treatment data, and genomic information all carry specific legal restrictions that vary by jurisdiction. A harmonization pipeline that ignores these boundaries may produce technically clean data that is legally unusable. In the United States, the Health Insurance Portability and Accountability Act (HIPAA) sets the baseline rules for how Protected Health Information (PHI) must be handled, covering everything from access controls and encryption to breach notification and minimum necessary use. Any harmonization pipeline that touches patient data, and any MCP endpoint or AI agent that consumes it, must operate within HIPAA's Privacy and Security Rules. This isn't a separate work-stream from harmonization; it's the regulatory floor that every architectural decision must respect from day one.
Auditability means maintaining a clear trail of every transformation, access, and inference. When an AI agent recommends a clinical action, the organization needs to trace the chain from source data through harmonization through the agent's reasoning, both for clinical accountability and regulatory compliance.
These concerns don't replace the technical harmonization work described above, they sit on top of it. But any organization building toward AI-driven clinical workflows will find that governance questions surface quickly, and having provenance and trust baked into the canonical data layer from the start is far easier than retrofitting it later.
The Real Starting Line
There's a common misconception that AI in healthcare is primarily a model problem, that the path forward is about better algorithms, larger training sets, or more powerful LLMs. In practice, the organizations making real progress have learned the opposite lesson: the bottleneck is almost always the data.
A brilliant model reasoning on fragmented, ambiguously coded, inconsistently structured clinical data will produce confident nonsense. A modest model reasoning on harmonized, semantically rich, well-governed data will produce something a clinician can actually trust.
The standards exist. FHIR, SNOMED CT, LOINC, RxNorm, OMOP, they're mature, widely adopted, and battle-tested. The connectivity patterns are emerging, with MCP offering a plausible path toward standardized agentic access. The governance frameworks, HIPAA, provenance, consent, are well understood even if they're not always well implemented.
What most organizations are missing isn't technology. It's the disciplined, sustained investment in treating data harmonization not as a backend plumbing project, but as the strategic foundation everything else depends on. The organizations that make that investment now won't just be ready for today's AI applications. They'll be ready for the ones that haven't been invented yet.